Generally, polynomial algorithms in the sofia_redux.toolkit.fitting module
either fit polynomial coefficients to a sample distribution, or evaluate
polynomial coefficients at a given location. The polyfitnd() function
is used to derive polynomial coefficients, while poly1d() or
polynd() functions evaluate coefficients in 1 or N dimensions
respectively. The Polyfit class is capable of both deriving and
evaluating polynomials. However, if one only wishes to derive and then
evaluate a polynomial fit to data (i.e. resample) but does not require the
actual coefficients using piecewise fits, it is recommended to instead refer
to sofia_redux.toolkit.resampling, developed for just that purpose.
Polynomial Coefficients¶
In this scheme, coefficients are stored as an N-dimensional array with the size of each dimension equal to the polynomial degree (order + 1) set for that dimension. For example, if we wish to fit a set of 2-dimensional data with a 2nd order (degree=3) polynomial in x, and a 3rd order (degree=4) polynomial in y, an array of shape (3, 4) would be used to contain the polynomial coefficients necessary to model the data with the desired polynomial.
Polynomial functions model distribution \(x\) as:
\[f(x) = \sum_{p_{1}=0}^{o_{1}} \sum_{p_{2}=0}^{o_{2}} ... \sum_{p_{N}=0}^{o_{N}} a_{(p_{1},p_{2},...,p_{N})} x_{1}^{p_{1}} x_{2}^{p_{2}} ... x_{N}^{p_{N}}\]
in \(N\) dimensions where \(o_{i}\) is the polynomial order fitted to dimension \(i\), and \(a\) are the polynomial coefficients. For example, polynomial coefficients could be used to model the function
\[f(x) = 5 + 2x + 3xy + 0.5y^{2}\]
as
\[\begin{split}a = \begin{bmatrix} 5 & 2 & 0 \\ 0 & 3 & 0 \\ 0.5 & 0 & 0 \end{bmatrix}\end{split}\]
using a second order polynomial in both x and y.
The following example uses polyfitnd() to derive these coefficients:
from sofia_redux.toolkit.fitting.polynomial import polyfitnd
import numpy as np
y, x = np.mgrid[:5, :5]
z = 5 + (2 * x) + (3 * x * y) + (0.5 * y ** 2)
a = polyfitnd(x, y, z, 2)
assert np.allclose(a, [[5, 2, 0],
[0, 3, 0],
[0.5, 0, 0]])
These coefficients can then be evaluated using polynd():
from sofia_redux.toolkit.fitting.polynomial import polynd
fitted = polynd(np.stack((y, x)), a)
assert np.allclose(z, fitted)
Both derivation and evaluation can be performed by the Polyfit class:
from sofia_redux.toolkit.fitting.polynomial import Polyfit
pfit = Polyfit(x, y, z, 2)
assert np.allclose(pfit.get_coefficients(), a)
assert np.allclose(pfit(x, y), z)
Please note however, that in all the above examples, we have used the default redundant polynomial representation such that no coefficients will be calculated for \(a\) when \(\sum_{i=1}^{N}{p_i} > max(o)\). So for a redundant (default) polynomial fit of order 2 in both dimensions
\[\begin{split}a = \begin{bmatrix} 5 & 2 & 0 \\ 0 & 3 & - \\ 0.5 & - & - \end{bmatrix}\end{split}\]
where values marked with \(-\) will always be set to zero since they are defined as redundant.
This default redundancy behaviour may be overridden by explicitly providing
terms for which coefficients must be derived. The format of these terms
should be provided as an integer array of shape (n_terms, n_dimensions), with
each value corresponding to the power of a single term in a specific dimension.
For example, in 3-dimensions, the value [1, 2, 3] would result in a coefficient
being calculated for term \(xy^2z^3\). The
sofia_redux.toolkit.utilities.func.polyexp() may be used for the purposes of
creating either full or redundant set of terms.
from sofia_redux.toolkit.fitting.polynomial import polyexp
# Create a redundant set of terms
redundant = polyexp(2, ndim=2)
print(redundant)
# [[0 0]
# [1 0]
# [2 0]
# [0 1]
# [1 1]
# [0 2]]
The above redundant set results in a function of the form:
\[f(x) = a_{0,0} + a_{1,0}x + a_{2,0}x^2 + a_{0,1}y + a_{1,1}xy + a_{0,2}y^2\]
# Create a full set of terms
full = polyexp([2, 2])
print(full)
# [[0 0]
# [1 0]
# [2 0]
# [0 1]
# [1 1]
# [2 1]
# [0 2]
# [1 2]
# [2 2]]
The above full set results in a function of the form:
\[f(x) = a_{0,0} + a_{1,0}x + a_{2,0}x^2 + a_{0,1}y + a_{1,1}xy + a_{2,1}x^2y + a_{0,2}y^2 + a_{1,2}xy^2 + a_{2,2}x^2y^2\]
Alternatively, terms may be explicitly defined by the user:
user = [[2, 0], [1, 1]]
In this case, the user has specified that coefficients should only by derived for the terms in the following function:
\[f(x) = a_{2,0}x^2 + a_{1,1}xy\]
For example, attempting to fit the function
\[f(x) = 1 + 2xy + 0.1x^2y^2\]
can only be done with either a full or user defined set of terms since \(x^2y^2\) will never appear in a redundant set of a 2nd order 2-dimensional polynomial function:
y, x = np.mgrid[:5, :5]
z = 1 + (2 * x * y) + (0.1 * x ** 2 * y ** 2)
redundant_pfit = Polyfit(x, y, z, 2)
print(redundant_pfit.get_coefficients().round(decimals=2))
# [[ 3.8 -3.2 0.6]
# [-3.2 3.6 0. ]
# [ 0.6 0. 0. ]]
full_pfit = Polyfit(x, y, z, polyexp([2, 2]), set_exponents=True)
print(full_pfit.get_coefficients().round(decimals=2))
# [[ 1. -0. 0. ]
# [-0. 2. -0. ]
# [ 0. -0. 0.1]]
user_pfit = Polyfit(x, y, z, [[0, 0], [1, 1], [2, 2]], set_exponents=True)
print(user_pfit.get_coefficients().round(decimals=2))
# [[1. 0. 0. ]
# [0. 2. 0. ]
# [0. 0. 0.1]]
As can be seen above, the default redundant set of terms failed to derive the
correct coefficients, but made its best attempt using those terms available.
The full and user defined sets both returned the correct values. Note that in
order to override the default redundant behaviour, set_exponents=True must
be set as a keyword argument. Also note that while the redundant fit failed,
the resulting fit is the “best fit” with the terms available as can be
seen below:
import matplotlib.pyplot as plt
import numpy as np
from sofia_redux.toolkit.fitting.polynomial import Polyfit
y, x = np.mgrid[:5, :5]
z = 1 + (2 * x * y) + (0.1 * x ** 2 * y ** 2)
redundant_pfit = Polyfit(x, y, z, 2)
plt.figure(figsize=(5, 5))
plt.plot(z, label='Original')
plt.plot(redundant_pfit(x, y), '--', label="Redundant fit")
plt.xlabel('x')
plt.ylabel('f(x, y)')
plt.title("Redundant term set fit")
plt.legend()
(Source code, png, hires.png, pdf)
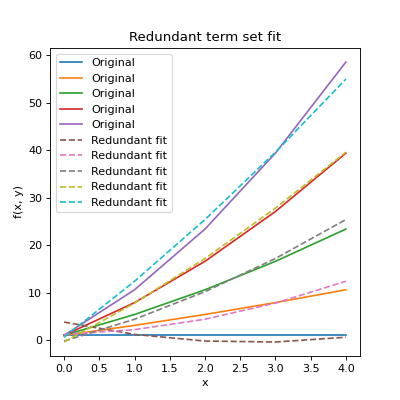
Plots for the full and user defined sets would reproduce the original data.
Fit Statistics¶
In addition to performing and evaluating a polynomial fit, Polyfit
can also return useful statistics including covariance terms. The
Polyfit.stats attribute is a namespace containing the following
statistics:
n: Number of samples
dof: Degrees of Freedom
fit: An array containing the fit to the original data
residuals: An array containing the difference between the data and the fit
sigma: An array containing the error of each polynomial coefficient
rms: The root-mean-square error on the fit
chi2: The \(\chi^2\) statistic on the fit
rchi2: The reduced \(\chi^2\) statistic on the fit
q: Goodness of fit, or survival function. The probability (0->1) thatone of the samples is greater than \(\chi^2\) away from the fit.
These can be accessed directly or printed to screen by applying the standard
Python print() function to an initialized Polyfit object. For
example:
import numpy as np
import matplotlib.pyplot as plt
from sofia_redux.toolkit.fitting.polynomial import Polyfit
# Create function
x = np.linspace(0, 2, 256)
y = -0.5 + (x - 2) ** 2
# add noise with a normal distribution
rand = np.random.RandomState(42)
noise = rand.normal(loc=0, scale=0.5, size=x.size)
y += noise
# Fit a 2nd order polynomial to the noisy data
# Since we know the scale of the error, it may be included
pfit = Polyfit(x, y, 2, error=0.5)
fig, ax = plt.subplots(nrows=1, ncols=2, figsize=(10, 5))
fig.tight_layout()
ax[0].plot(x, y, '.', label="Samples")
ax[0].plot(x, pfit.stats.fit, '-', label="Fit")
ax[0].legend()
ax[0].set_title("Polynomial fit to noisy data")
ax[1].plot(x, pfit.stats.residuals, '.')
ax[1].set_title("Residuals to the fit")
print(pfit)
(Source code, png, hires.png, pdf)
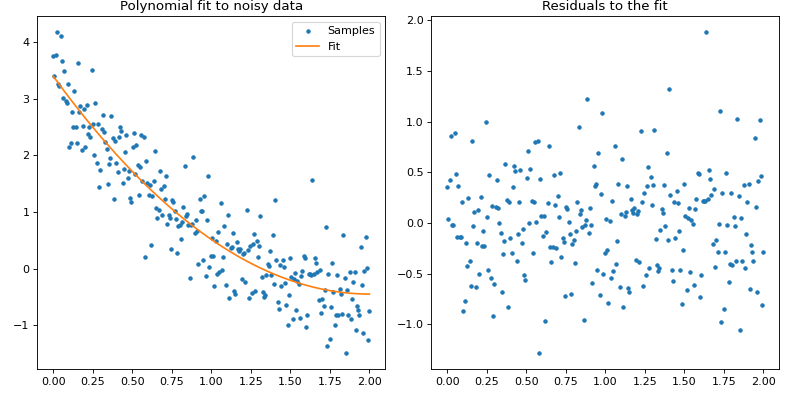
Textual output:
Name: Polyfit
Statistics
--------------------------------
Number of original points : 256
Number of NaNs : 0
Number of outliers : 0
Number of points fit : 256
Degrees of freedom : 253
Chi-Squared : 240.743406
Reduced Chi-Squared : 0.951555
Goodness-of-fit (Q) : 0.300082
RMS deviation of fit : 0.485822
Exponents : Coefficients
--------------------------------
(0,) : 3.396036 +/- 0.093022
(1,) : -3.840006 +/- 0.214879
(2,) : 0.958356 +/- 0.104002
Covariance¶
The full covariance matrix is available in the Polyfit.covariance
attribute. Note that this is an (n_coefficient, n_coefficient) giving the
covariance between terms, with each axis ordered in the same manner as the
terms. The order of the terms can be determined from the
Polyfit.exponents attribute. For example:
import numpy as np
from sofia_redux.toolkit.fitting.polynomial import Polyfit
# Create a function
y, x = np.mgrid[:10, :10]
z = 1 - (5 * x) + (3 * y)
pfit = Polyfit(x, y, z, 1)
print("Exponents:\n%s\n" % pfit.exponents)
print("Covariance:\n%s" % pfit.covariance)
Output:
Exponents:
[[0 0]
[1 0]
[0 1]]
Covariance:
[[ 5.90909091e-02 -5.45454545e-03 -5.45454545e-03]
[-5.45454545e-03 1.21212121e-03 4.33680869e-19]
[-5.45454545e-03 -0.00000000e+00 1.21212121e-03]]
In the above example, the rows and columns of the covariance matrix are ordered as \(a_{0,0}, a_{1, 0}, a_{0,1}\) for the terms \(x^0y^0, x, y\).
Disabling Statistics¶
If statistics are not required, there is no need for a covariance matrix, and
processing speed is important, statistical calculations may be disabled by
setting stats=False and/or covar=False during intialization. If statistics
are disabled, they will no longer be calculated or displayed. If covariance
calculations are disabled, the Polyfit.covariance attribute will be
set to None.
import numpy as np
from sofia_redux.toolkit.fitting.polynomial import Polyfit
y, x = np.mgrid[:10, :10]
z = 1 - (5 * x) + (3 * y)
pfit = Polyfit(x, y, z, 1, stats=False, covar=False)
assert pfit.covariance is None
print(pfit)
Output:
Name: Polyfit
Exponents : Coefficients
--------------------------------
(0, 0) : 1.000000
(1, 0) : -5.000000
(0, 1) : 3.000000
However, if robust outlier rejection is enabled, by necessity, both statistics and covariance will be calculated regardless of user wishes.
Robust Outlier Rejection¶
Robust outlier rejection may be enabled by setting robust > 0 during
Polyfit initialization. If set, outliers are identified through an
iterative process: fitting the data then excluding any samples that fall
outside robust sigmas from the fit before repeating. Iterations are
terminated after:
A certain number of iterations have occurred.
The relative delta between successive residual RMS values is less than a set value.
Too many samples are excluded such that fitting the desired order of polynomial will not be possible.
The fit failed.
Whether the iteration succeeded can be determined from the
Polyfit.success attribute, while the termination condition encountered
is in the Polyfit.termination attribute. Which samples were excluded
during the rejection process can be determined from the Polyfit.mask
attribute. Additional statistics will also be displayed if statistics are
printed to screen. For example:
import numpy as np
from sofia_redux.toolkit.fitting.polynomial import Polyfit
# Create a function
x = np.linspace(0, 5, 256)
y = 1 + x
rand = np.random.RandomState(42)
# Add noise
noise = rand.normal(loc=0, scale=0.5, size=y.shape)
y += noise
# Add some obvious outliers, throwing off the fit
inds = rand.randint(0, 255, 25)
y[inds] += 5
# Standard fit
pfit = Polyfit(x, y, 1)
# Robust fit with 3 sigma outlier rejection
rfit = Polyfit(x, y, 1, robust=3)
outliers = np.argwhere(~rfit.mask)[:, 0]
plt.figure(figsize=(5, 5))
plt.plot(x, y, '.', label='Samples')
plt.plot(x, pfit.stats.fit, label='Standard Fit')
plt.plot(x, rfit.stats.fit, label='Robust Fit')
plt.plot(x[outliers], y[outliers], 'x', color='red', label='Outliers')
plt.title("Robust Outlier Rejection")
plt.xlabel("x")
plt.ylabel("f(x)")
plt.legend()
# Display robust statistics
print(rfit)
(Source code, png, hires.png, pdf)
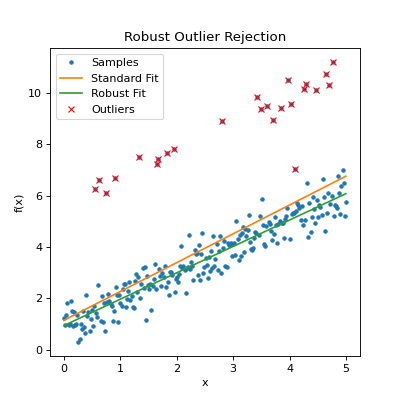
Output:
Name: Polyfit
Statistics
--------------------------------
Number of origial points : 256
Number of NaNs : 0
Number of outliers : 24
Number of points fit : 232
Degrees of freedom : 230
Chi-Squared : 53.825532
Reduced Chi-Squared : 0.234024
Goodness-of-fit (Q) : 0.000000
RMS deviation of fit : 0.482712
Outlier sigma threshold : 3
eps (delta_sigma/sigma) : 0.01
Iterations : 3
Iteration termination : delta_rms = 0
Exponents : Coefficients
--------------------------------
(0,) : 0.941421 +/- 0.129497
(1,) : 1.026215 +/- 0.045540
The above report indicates 24 outliers were found in which the residual was greater than the \(3\sigma\) limit. 3 iterations were required, terminated when no further outliers could be found (delta_rms = 0).
Special Case 1-D and 2-D Functions¶
For quick and easy derivation and evaluation of polynomial coefficients in one or two dimensions, several functions are available that deviate from the more general API presented above.
1-Dimensional Polynomial Derivation and Evaluation¶
For fitting a 1-dimensional polynomial to data where no special considerations
are necessary, there are already many functions available in standard
Python packages such as numpy.polyfit(), so no attempt has been made to
recreate another version. polyfit() is perfectly capable of handling 1
dimensional cases with the added benefits of
sofia_redux.toolkit.fitting.polynomial.Polyfit functionality.
However, for 1-dimensional evaluation, poly1d() is available to
evaluate coefficients, and optional variance if either the full covariance
matrix is provided, or the diagonal of the covariance matrix is given.
import numpy as np
import matplotlib.pyplot as plt
from sofia_redux.toolkit.fitting.polynomial import poly1d, polyfitnd
x = np.linspace(0, 4 * np.pi, 1000)
y = np.sin(x)
error = np.random.RandomState(41).normal(loc=0, scale=0.1, size=x.size)
y += error
# use polyfitnd to fit a polynomial and get the covariance on the fit
# coefficients
coeffs, cvar = polyfitnd(x, y, 7, covar=True)
yfit, yvar = poly1d(x, coeffs, covar=cvar)
error = np.sqrt(yvar)
plt.figure(figsize=(7, 5))
plt.plot(x, y, '.', markersize=3, label='data', color='blue')
plt.fill_between(x, yfit - error, yfit + error, color='red',
label=r'$1\sigma$ fit error')
plt.plot(x, yfit, label='fit', color='lime')
plt.legend(loc='lower left')
plt.title("7th order polynomial fit and fit error")
plt.xlabel('x')
plt.ylabel('y')
(Source code, png, hires.png, pdf)
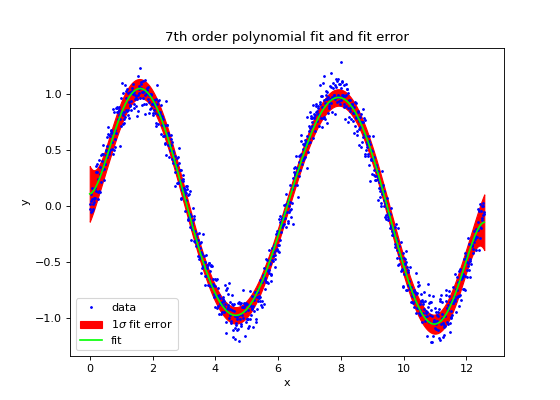
Please note that polynd() and Polyfit are both capable of
producing the same results as poly1d(). However, poly1d() is
a lighter version using numpy.poly1d() to evaluate coefficients rather
than the engine used by polynd().
2-Dimensional Polynomial Derivation and Evaluation¶
For 2-dimensional data, the polyfit2d() and poly2d() distinguish
between the full set of polynomial coefficients, and the more robust
redundancy controlled term coefficients in the same way as polyfitnd().
This distinction is controlled by the full keyword. The following table
gives an example of the coefficients for a 2nd order polynomial with
full=True and full=False. Note that the order of coefficients in the
output table is different from polyfitnd(), with x coefficients running
along columns and y coefficients by rows, where the exponent of the coefficient
term is given by it’s index. The order of polynomial must be given separately
for the x and y directions via the kx and ky keyword arguments (default=2).
x0 |
x1 |
x2 |
|
y0 |
c0,0 |
c1,0 |
c2,0 |
y1 |
c0,1 |
c1,1 |
c2,1 |
y2 |
c0,2 |
c1,2 |
c2,2 |
x0 |
x1 |
x2 |
|
y0 |
c0,0 |
c1,0 |
c2,0 |
y1 |
c0,1 |
c1,1 |
|
y2 |
c0,2 |
2-Dimensional Examples¶
polyfit2d()derives polynomial coefficients describing a surface:>>> import numpy as np >>> from sofia_redux.toolkit.fitting.polynomial import polyfit2d >>> y, x = np.mgrid[:5, :5] >>> z = 0.5 + (0.4 * x) + (0.3 * y) + (0.2 * x * y) + (0.1 * x ** 2) >>> c = polyfit2d(x, y, z, kx=2, ky=2) >>> print(np.abs(np.round(c, decimals=3))) [[0.5 0.4 0.1] [0.3 0.2 0. ] [0. 0. 0. ]]
poly2d()evaluates 2-dimensional polynomial coefficients:>>> from sofia_redux.toolkit.fitting.polynomial import poly2d >>> np.allclose(poly2d(x, y, c), z) True
polyinterp2d()derivates and evaluates 2-D polynomial coefficients:>>> from sofia_redux.toolkit.fitting.polynomial import polyinterp2d >>> np.allclose(polyinterp2d(x, y, z, x, y, ky=2, kx=2), z) True
These 2-dimensional functions are designed as lighter versions of
polyfitnd() and polynd(), so should fit and evaluate data slightly
faster.