The sofia_redux.toolkit.convolve.kernel module contains the
ConvolveBase class, inheritors of ConvolveBase, and
functional wrappers for these classes.
Convolution¶
Currently the following classes may be used to filter and smooth data via kernel convolution in N-dimensions:
BoxConvolve
KernelConvolve
SavgolConvolve
All classes inherit from the sofia_redux.toolkit.base.Model class, shared
by sofia_redux.toolkit.fitting.polynomial.Polyfit, so much of the statistical
functionality and robust outlier rejection will also apply in the following
section. While there are many, many algorithms available for convolution in
the Python ecosystem already, the main functionality of these classes is that
they allow for:
N-dimensional
Interpolation over masked/NaN values
Error propagation
Statistical analysis and outlier rejection
This is achieved using a combination of linear interpolation with Delaunay triangulation (when more than a single dimension is involved). Therefore, if none of the above features are required, it is recommended to use one of the more standard algorithms for speed by avoiding the triangulation step. Regardless, these algorithms are still quite fast.
Basic Functionality¶
import numpy as np
import matplotlib.pyplot as plt
import imageio
from sofia_redux.toolkit.convolve.kernel import BoxConvolve
image = imageio.imread('imageio:coffee.png').sum(axis=2) # Gray scale
image = (image - image.min()) / (np.ptp(image)) # normalize for plotting
mean_smooth = BoxConvolve(image, 11) # an 11 x 11 box filter
fig, ax = plt.subplots(nrows=1, ncols=2, figsize=(10, 5))
fig.tight_layout()
ax[0].imshow(image, cmap='gray')
ax[0].set_title("Original Image")
ax[1].imshow(mean_smooth.result, cmap='gray')
ax[1].set_title("Smoothed Image")
(Source code, png, hires.png, pdf)
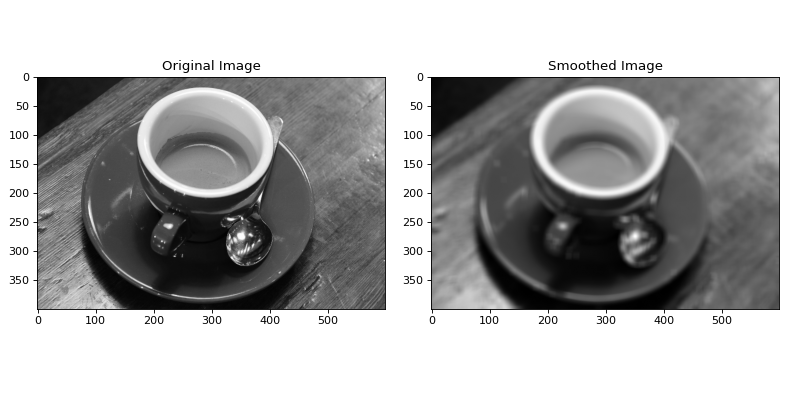

In the above example, BoxConvolve was used to set the value of each
pixel to the mean value of an (11, 11) box kernel centered on each pixel.
BoxConvolve is convenient, but only really just a child of
KernelConvolve which does the same thing except allows the user to
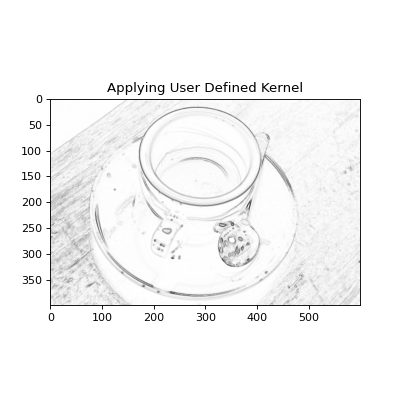
define their own smoothing kernel. For example, we could create a user
defined Sobel filter to enhance edges in the image:
import numpy as np
import matplotlib.pyplot as plt
import imageio
from sofia_redux.toolkit.convolve.kernel import KernelConvolve
image = imageio.imread('imageio:coffee.png').sum(axis=2) # Gray scale
image = (image - image.min()) / (np.ptp(image)) # normalize for plotting
# Create a Sobel filter
sobel = np.array([[-1, 0, 1], [-2, 0, 2], [-1, 0, 1.]])
# Edges in 1 direction
sx = KernelConvolve(image, sobel, normalize=False).result
# Edges in the other
sy = KernelConvolve(image, sobel.T, normalize=False).result
# Edge amplitude
sxy = np.hypot(sx, sy)
plt.figure(figsize=(5, 5))
plt.imshow(sxy, cmap='gray_r')
plt.title("Applying User Defined Kernel")
(Source code, png, hires.png, pdf)
Normalization¶
By default, a kernel (\(K\)) will be normalized before being applied to the data such that
However, this will only be done if \(\sum{K} \neq 0\). To keep the
original \(K\), rather the normalized kernel \(\hat{K}\), set
normalize=False during initialization.
s = KernelConvolve(data, kernel, normalize=False)
Masked Values, NaN handling and Error Propagation¶
Missing (NaN) values and masked values (those defined by the user to ignore)
are treated in the same way by the ConvolveBase class. Linear
interpolation is first performed to generate values at these locations before
convolution.
Errors are propagated, consistent with the initial interpolation and subsequent convolution. For 1-dimensional data, standard linear interpolation is performed. For higher dimensions, Delaunay triangulation is used instead. Therefore, one would expect 3 points used for the calculation of each interpolant in 2 dimensions, 4 points in 3-dimensions etc. This is reflected during error propagation which will be contingent on the number of points and structure of the tessellation.
In a similar way to most other classes in sofia_redux.toolkit.convolve, NaNs
may be ignored (default), or propagated (ignorenans=False). Note however,
that NaNs will then propagate through the convolution operation, expanding
any NaN value through the entirety of the kernel. I.e., if a single NaN is
present in the original data and convolved with a kernel of width 5, a NaN
region of width=5 will be present in the output convolution.
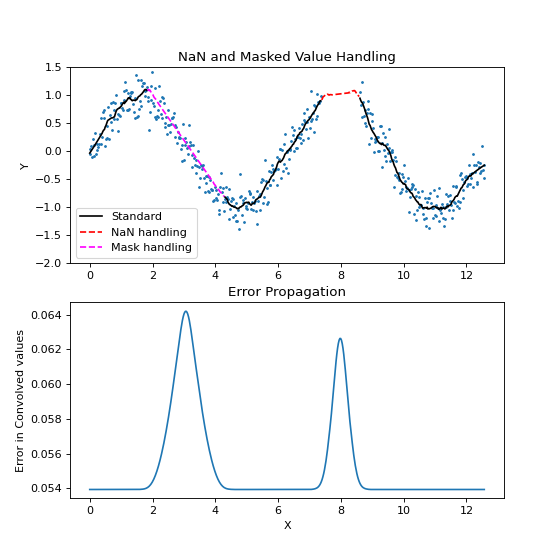
The example below uses a Savitzky-Golay filter which approximates polynomial interpolation for regularly spaced data. Large regions of the samples are set to NaN values or masked out.
import numpy as np
import matplotlib.pyplot as plt
from sofia_redux.toolkit.convolve.kernel import SavgolConvolve
x = np.linspace(0, 4 * np.pi, 512)
y = np.sin(x)
error = 0.2
rand = np.random.RandomState(41)
noise = rand.normal(loc=0.0, scale=error, size=x.size)
y += noise
# Add NaN Values
y[300:350] = np.nan
# Mask certain values to be excluded from fit, but interpolated over
mask = np.full(x.size, True)
mask[75:175] = False
width = 31
s = SavgolConvolve(x, y, width, error=error, mask=mask, order=2)
fig, ax = plt.subplots(nrows=2, ncols=1, figsize=(7, 7))
black_plot = s.result.copy()
black_plot[~mask] = np.nan
black_plot[np.isnan(y)] = np.nan
ax[0].set_title("NaN and Masked Value Handling")
ax[0].plot(x, y, '.', markersize=3)
ax[0].plot(x, black_plot, color='black',
label='Standard')
ax[0].plot(x[300:350], s.result[300:350], '--', color='red',
label='NaN handling')
ax[0].plot(x[75:175], s.result[75:175], '--', color='magenta',
label='Mask handling')
ax[0].legend(loc='lower left')
ax[0].set_ylim(-2, 1.5)
ax[1].set_title("Error Propagation")
ax[1].plot(x, s.error)
ax[1].set_xlabel("X")
ax[1].set_ylabel("Error in Convolved values")
ax[0].set_ylabel("Y")
(Source code, png, hires.png, pdf)
Robust Outlier Rejection and Statistics for the Convolve Class¶
In a similar way to the sofia_redux.toolkit.fitting.polynomial.Polyfit class,
children of Convolve may perform robust outlier rejection. An outlier
is determined from the magnitude of the residual of the original data samples
to the convolved values. Such outliers will be masked out and replaced with a
linear interpolation from values determined to be within the rejection
threshold. This process is repeated until no new outliers are found or one of
the other conditions described in robust outlier rejection for
sofia_redux.toolkit.fitting.polynomial.Polyfit is met.
import imageio
import numpy as np
import matplotlib.pyplot as plt
from sofia_redux.toolkit.convolve.kernel import SavgolConvolve
# Normalize for plotting
image = imageio.imread('imageio:immunohistochemistry.png').astype(float)
image = image.sum(axis=-1)
image -= image.min()
image /= image.max()
# Add some bad values
rand = np.random.RandomState(41)
inds = rand.choice(image.shape[0], 100), rand.choice(image.shape[1], 100)
image[inds] += 2
mask = np.full(image.shape, True)
mask[inds] = False
s = SavgolConvolve(image, 7, order=3)
s_robust = SavgolConvolve(image, 7, robust=5, order=3)
fig, ax = plt.subplots(nrows=2, ncols=2, figsize=(10, 10))
ax[0, 0].imshow(image)
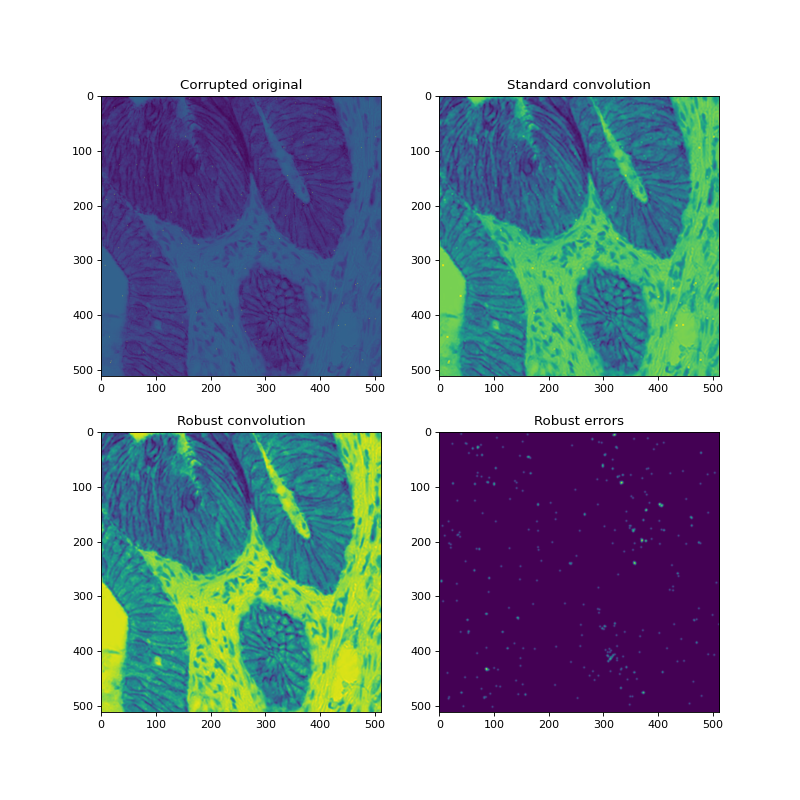
ax[0, 0].set_title("Corrupted original")
ax[0, 1].imshow(s.result)
ax[0, 1].set_title("Standard convolution")
ax[1, 0].imshow(s_robust.result)
ax[1, 0].set_title("Robust convolution")
ax[1, 1].imshow(s_robust.error)
ax[1, 1].set_title("Robust errors")
# Display statistics
print(s_robust)
(Source code, png, hires.png, pdf)
Output:
Name: SavgolConvolve
Statistics
--------------------------------
Number of original points : 262144
Number of NaNs : 0
Number of outliers : 420
Number of points fit : 261724
Degrees of freedom : 261724
Chi-Squared : 1841.870250
Reduced Chi-Squared : 0.007037
Goodness-of-fit (Q) : 0.000000
RMS deviation of fit : 0.028076
Outlier sigma threshold : 5
eps (delta_sigma/sigma) : 0.01
Iterations : 3
Iteration termination : delta_rms/rms = 0.001941
The above example shows an artificially corrupted image, the effects of convolution without removing outliers, a convolution with outliers removed, and the resulting errors from the robust convolution algorithm. Visually, all outliers have been removed.